News
Every dataset has a story - MedOBIS publishes its first micro-Computed Tomography Morphological dataset
26 March 2026
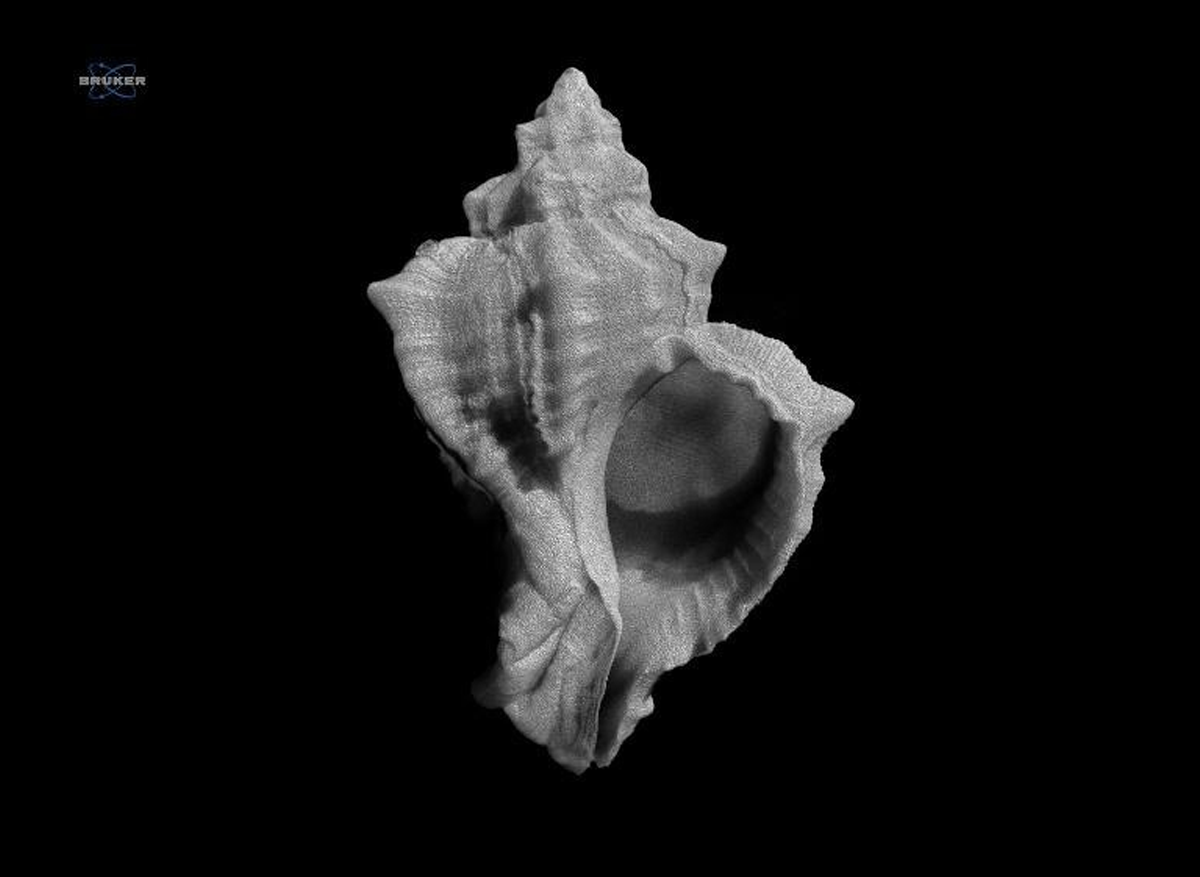

A micro-CT scan of Hexaplex trunculus, the banded dye-murex, one of two species at the centre of MedOBIS's innovative morphological dataset.
Emerging data types often pose challenges for repositories in terms of management, standardization, and integration. MedOBIS, the OBIS Node for the Mediterranean, proposed a solution for this challenge and published its first-ever Darwin Core-formatted dataset that included micro-Computed Tomography Morphological image-derived information, along with biodiversity and environmental data on two species, Chondrilla nucula and Hexaplex trunculus.
The MACCIMO project—Multi-level Approaches to Assess Climate Change Impact on Marine Organisms—investigates how climate change affects sessile marine invertebrates through a multipronged strategy that integrates multiple scientific approaches. This project was funded by the Hellenic Foundation for Research and Innovation (HFRI) under the “2nd Call for HFRI Research Projects to support Faculty Members & Researchers” (Project Number: 3280). Sessile marine invertebrates are particularly sensitive to climate-related stressors due to their limited mobility and their inability to escape adverse environmental conditions. As part of experimentally simulated climate change scenarios, one of the project’s aims was to investigate potential morphological changes in two species: the chicken-liver sponge (Chondrilla nucula, a sessile sponge permanently attached to the substrate) and the banded dye-murex (Hexaplex trunculus, a medium-sized, low mobility sea snail historically famous for its use in creating purple and blue dyes). By examining how the morphology of these species responds to simulated climate stressors, the project aims to identify traits that can serve as indicators of environmental change.
To conduct these investigations, the project’s team performed three-dimensional analyses using micro-Computed Tomographic (micro-CT) scans. Microtomography is a non-destructive imaging technique based on X-Rays, allowing the creation of high-resolution three-dimensional data. Once generated, these 3D datasets had to be prepared and organised for publication. Making such observations available as open, interoperable data upgrades their actionability: standardised 3D morphological records can be compared, combined with data from other institutions, and reused globally. “From the start, we aimed to make the image data of the project available and accessible to all,” says Dimitra Mavraki, MedOBIS Node Manager. The process was a first for the team and required creative thinking.
The MedOBIS team first focused on understanding micro-computed tomography and the data associated with that imaging technology. The team approached Darwin Core compliance using a hierarchical schema, starting with the sampling data and curation process data. “We used an Event core, where the parent event represents the sampling event, and a child event represents the documentation of specimens in the internal micro-CT library. Each specimen was then represented through an Occurrence extension linked to these events,” says Dimitra Mavraki. She goes on to explain that the team’s goal was to publish the quantitative outputs derived from micro-CT analyses alongside the biological occurrences, so they had to capture the micro-CT–derived parameters using the extended MeasurementOrFact (eMoF) extension. Using the eMoF extension allowed them to express each derived measurement in a standard, machine-readable manner while remaining within Darwin Core standards.
Additionally, the team mobilized persistent internal micro-CT library identifiers to maintain traceability and consistency across systems, where the parentEventID reflects the event type “Sample Documentation”, while the eventID reflects the scan event type “MicroCT_Scan.” Each ID is included in both the eventID and occurrenceID names. Finally, the MedOBIS team applied the OBIS Quality Assessment and Quality Control steps before publishing.
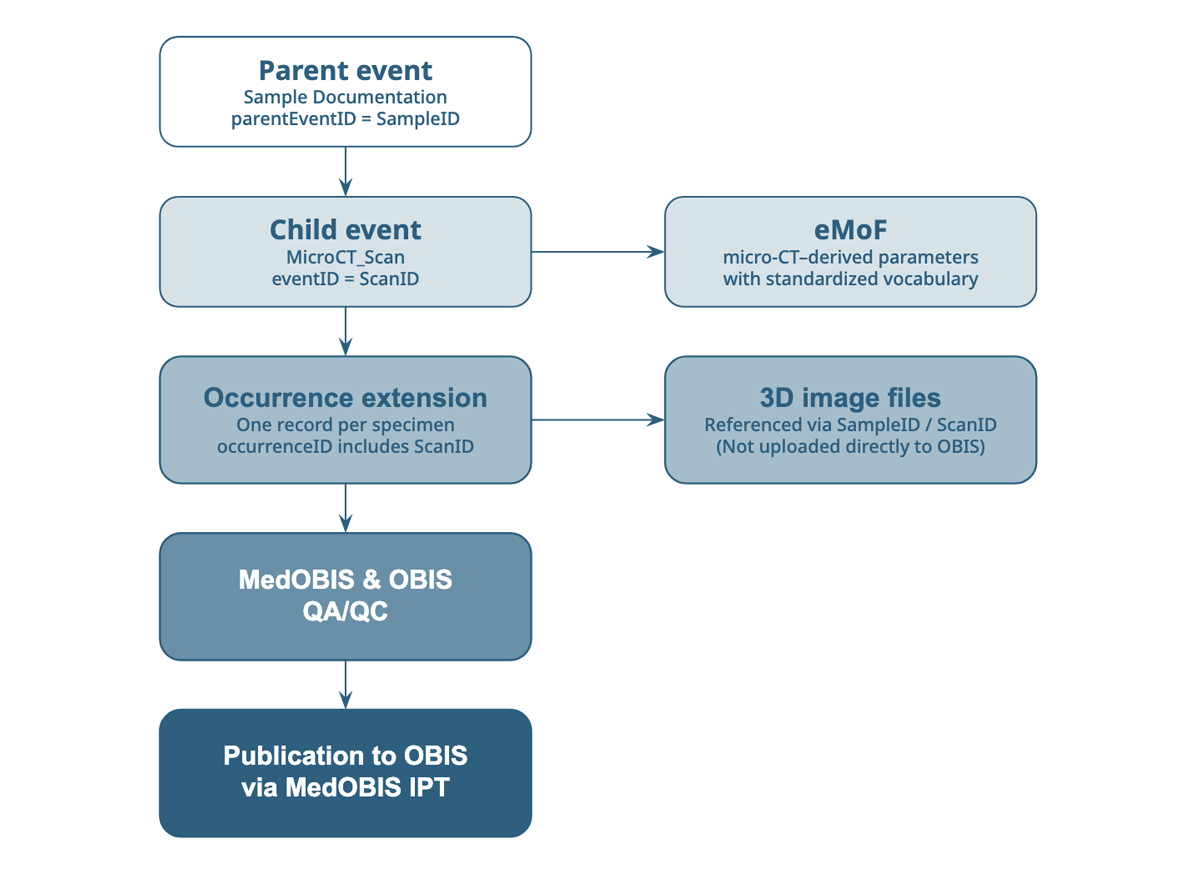
Overview of the event hierarchy through the publishing process.
Working hands-on with micro-CT data helped MedOBIS to highlight several challenges in data curation and management. Unlike with biodiversity data, micro-CT datasets still lack widely adopted metadata standards. In addition, their large file sizes make data organization and long-term storage more demanding. For this reason, the actual micro-CT image files are not uploaded directly to OBIS. Instead, within the OBIS dataset, these 3D resources are referenced by maintaining the relevant micro-CT identifiers (SampleID/ScanID) in the eventID field to ensure consistency at this stage. The long-term objective of MedOBIS is to link these 3D resources directly through their URLs.
3D volume rendering of Marphysa victori
Yet, this milestone felt like a natural step forward. “Micro-CT technology has been used at the Hellenic Centre for Marine Research, the Institution hosting MedOBIS, for many years,” says Dimitra Mavraki. “We knew that the technology had the potential to enrich the datasets we publish, and create new opportunities for research, comparison, education, and future technological applications”. She explains that until now, such scans were hosted in a dedicated online platform featuring a collection of annotated 3D specimens that users can explore interactively. Bringing these resources to MedOBIS opens new possibilities for connecting morphological research with global biodiversity data networks and sharing it with the world.
The team sees this first publication as a proof-of-concept for micro-CT specimen imaging within OBIS, successfully demonstrating that imaging-derived outputs can be standardized and made FAIR. Their new objective? Automate and scale the process to publish into OBIS all the specimens entering the micro-CT library of the Hellenic Centre for Marine Research. When asked if she has advice for OBIS Nodes who want to explore new data formats, Dimitra has no hesitation: “Explore new data types, even if stepping into unfamiliar territory can be challenging. It is incredibly rewarding. Working with new kinds of data pushes us to think differently, collaborate more closely, and develop solutions that benefit the wider scientific community. In the end, these efforts not only advance our infrastructures but also make our work more meaningful and impactful,” she concludes. ◼️

